Genetic Mutations in Autosomal Dominant Polycystic Kidney Disease Type-1
Chandra Sanwal, Karim Nooruddin, Chirag Patel, Tiannan Zhang, Anita Bhagavathula, Steven Gates and Antonio Guzman
DOI10.21767/2472-5056.100074
Department of Internal Medicine, Corpus Christi Medical Center, USA
- *Corresponding Author:
- Chandra Sanwal
Department of Internal Medicine
Corpus Christi Medical Center, USA
Tel: 1-813-5557777
E-mail: chandrasanwal7@gmail.com
Received date: February 02, 2019; Accepted date: February 18, 2019; Published date: February 21, 2019
Citation: Sanwal C, Nooruddin K, Patel C, Zhang T, Bhagavathula A, et al. (2019) Genetic Mutations in Autosomal Dominant Polycystic Kidney Disease Type-1. J Clin Exp Nephrol Vol.4 No.1:02. Doi: 10.21767/2472-5056.100074
Copyright: © 2019 Sanwal C, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Polycystic kidney disease (PKD) is mainly characterized by formation of cysts in kidney with associated kidney malfunction. The cysts eventually grow larger leading to end stage renal failure (ESRF) requiring kidney dialysis or transplant. There are various genetic forms of PKD: autosomal dominant type 1 (ADPKD1), type 2 (ADPKD2) and even a third type (ADPKD3) is known to exist. There is also a recessive form of PKD (ARPKD). The autosomal dominant type is caused due a single defective dominant gene in a single chromosome of the homologous pair among the autosomes and the recessive type is caused due to defect in both genes of a homologous pair. Each of these PKD types adversely impact structure and function of their associated protein products. For instance, ADPKD1 impacts polycystin-1, ADPKD2 impacts polycystin-2, and ARPKD impacts fibrocystin. There are several known mutations for each of these PKD disease types, e.g., there are over 250 known mutations of ADPKD1 gene. In this paper we restrict ourselves to ADPKD1 and a specific mutation (L845S) to demonstrate how a genetic mutation leads to malfunction of PKD1 gene.
Keywords
Polycystic kidney disease; Dialysis; Genetic mutations
Introduction
Among the different types of polycystic kidney diseases (PKD), Autosomal Polycystic Kidney Disease (ADPKD) is one of the most common disorders in humans and it impacts 1 out of 1000 individuals [1]. Apart from kidney malfunction due to cyst formation, it is also associated with cysts in liver, pancreas, spleen, kidneys, heart valve dysfunctions and brain aneurysms [2].
At the phenotypic level, all ADPKD types are indistinguishable, but at genetic level, two loci PKD1 and PKD2, have been identified [3]. The location of PKD1 is 16p13.3 (short of arm of chromosome 16) and that of PKD2 is 4q22.1 (long arm of chromosome 4) [3]. Mutations of PKD1 (ADPKD1) account for 90% of the cases of ADPKD. The location of ADPKD1 gene is proximal to that of Tuberculosis Sclerosis gene (TSC2) gene [3]. In fact, some TSC2 patients showed similar cysts as in ADPKD1, due to translocation of a long piece of the gene that included portions of both TSC2 and PKD1. This led researchers to the locus 16p13.3 and later it was confirmed that this location also produces a separate protein known as polycystin-1 (aka PKD1) [3].
It is observed in humans and rats that heterozygous mutations in either PKD1 or PKD2 show up typically near mid-life. In humans the disease shows up anytime between infancy and old age (80 years) [1]. Single homozygous mutations in either one of PKD1 or PKD2 genes, where both alleles on the homologous pair of chromosomes are mutated, or double heterozygous mutations in both PKD1 and PKD2, are lethal in very early embryonic stage [4].
A two-hit mechanism of development of the disease among the heterozygous ADPKD1 individuals is widely accepted [5,6]. Even though in early years, only one functional allele is sufficient to sustain the near normal kidney function, but as the individual reaches near mid-life, it is postulated that the single functional gene of the homologous pair becomes dysfunctional due to an associated somatic mutation. The trigger could be minor kidney damage such as a temporary obstructive or restriction to blood flow (ischemia) to renal epithelial cells, which initiates a repair mechanism, leading to a somatic mutation in the lone functional gene.
Apart from mutation defects, both up regulation and down regulation of PKD1 or PKD2 are known to cause cystic defects [7]. This has been demonstrated in experimental settings in transgenic murine cells. Evidently, there is a delicate balance between the activity of both PKD1 and PKD2 genes in a normal individual.
Both PKD1 and PKD2 are expressed in greater amounts in embryonic stages in kidney nephron development (nephrogenesis) than the adult tissues; PKD2 being expressed earlier and more diffusely than PKD1 [8]. The highest expression of PKD1 in adult tissues was seen in heart and brain. It was weaker in skeletal muscle and pancreas and weakest in the kidney.
PKD1 spans 52kb of genomic DNA, which is quite large. It has 46 exons, and its mRNA transcript is 14kb in length. The protein product of PKD1 has 4302 amino acids [8]. The exons of the gene show alternative splicing [3]. The protein is targeted to epithelial tissues of kidney tubules, liver and pancreatic ducts [9]. Even though the structure of PKD1 gene has been identified, there is it still a lack of comprehensive understanding of its various functions in a normal or diseased individual. The reason being it’s a large protein with multiple domains and plays a complicated role in various extracellular and intracellular functions, such as cell to cell communication or transcriptional control. There are large numbers of mutations in the PKD1 gene [10]. Some are even known to be private at the level of an individual or a family [2].
PKD1 is a well preserved gene and its homologues are found in the mouse, the Fugu fish, the sea urchin, and the worm Caenorhabditis elegans [11].
PKD1 structure
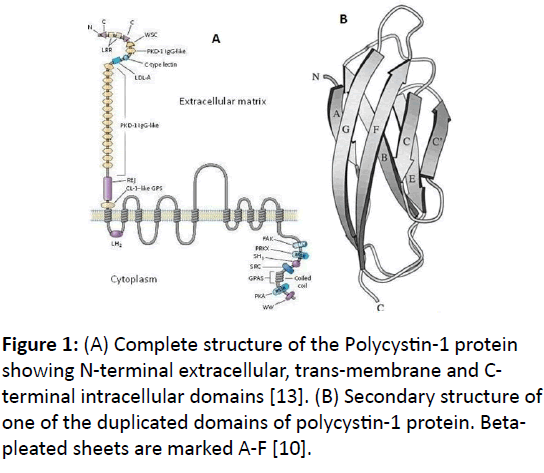
A high level structure of PKD1 protein starting from the Nterminus to C-terminus is described below [12]. It is a large integral membrane protein with 75% of the protein located in the extracellular region, 11 trans-membrane domains and 200 amino acids in the intracellular region (Figure 1A) [4].
Signal peptide
The protein starts with as a 23 amino acid signal sequence, which is found in other proteins [3]. Signal sequences direct the targeting of a protein to its final destination.
Leucine rich repeats (LRRs)
There are 2 LRR regions coming from exon 2 and 3, flanked with cysteine rich regions on N-terminus and C-terminus. LRRs are found in many other proteins and have various roles. They participate in protein-protein interaction by forming a horshoe like complex, such as the adhesive platelet glycoproteins [3]. There are other proteins which also have LRR regions flanked by cysteines, such as Toll, Slit etc., which relay signal to the cytoplasm [3].
C-type lectin domain
There is a single C-type (calcium dependent) lectin domain encoded by exons 6 and 7 [3]. This domain is believed to be involved in extracellular binding of carbohydrate residues to transfer glycosylated enzymes into the cell or participate in cell adhesion.
LDL-A domain
Downstream to the C-type lectin domain, encoded from exon 10, is a region consisting of about 40 amino acids, known as LDLA (lippoprotein A) domain. It is cysteine rich and hydrophobic [3]. This region is similar to regions found in LDL receptors and is believed to have similar function in ligand binding.
Duplicated PKD domains
There is a region of about 80 amino acids, which is duplicated in 14-16 copies across the length of PKD1 protein (Figure 1B) [3,8]. It makes 30% of the total amino acid count of the PKD1 protein. The first PKD1 domain is coded from exon 5 is located between LRRs and C-type lectin domain. The remaining 12-14 PKD domains are located contiguously downstream to LDL-A domain. Each of these PKD domains is organized as an antiparallel beta-pleated sandwich fold with a hydrophobic core [8] (Figure 1B). The PKD domains have immunoglobin (Ig) like folds and are believed to have similar function for form ligand-binding sites of surface proteins.
REJ domain
The Receptor for Egg Jelly (REJ) protein domain is similar to a protein found in sea urchin [13,14]. In sea urchin, the REJ is a membrane protein found in the egg that interacts with membrane glycoproteins of the sperm to control the acrosome reaction. Acrosome is the head part of a sperm and its reaction facilitates fusing of the sperm and the egg by breaking the barrier membranes of both egg and the sperm. Since acrosome reaction is assisted by release of Na+ and Ca2+ ions, the REJ domain in PKD1 is believed to regulate channel transport of these ions in epithelial cells.
GPS domain
The G-Protein Coupled Receptor (GPCR) Proteolytic Site (GPS) plays a role in post translational modification as described in the section below.
Intracellular domains
The intracellular LH2 domain is a site for possible interaction with lipids. Towards the C-terminal, the remaining 200 amino acid long part of PKD1 contains sites for protein-interaction and phosphorylation-signaling [13].
L845S gene mutation
This mutation is a missense mutation at amino acid position 845 in cDNA, where acid neutral and hydrophobic leucine is changed to polar serine. In reference to the genomic DNA, it is the position of the 6th amino acid starting from the N-terminal of PKD domain 2.
The exact DNA codon change is from TTG to TCG that corresponds to mRNA codon change of UUG (Leu) to UCG (Ser). This mutation was detected in a patient by loss of a site after cleavage by restriction enzyme StyI that did not match the cleavage of the normal gene [5].
Possible cause of L845S mutation
Apart from L845S, there are other various documented missense mutations in the PKD1 gene, e.g., R324L, W1399R, P1786L [2,5]. There are also nonsense (premature stop codon) and frame-shift (insertions and deletions) mutations. One study has attributed gene conversion as the most likely cause of such missense mutations [2,15]. Gene conversion is a non-reciprocal exchange of genetic material between the donor and recipient strands. Gene conversion is opposite of genetic recombination, where genetic material is exchanged in a reciprocal manner between homologous chromosomes. Watnick et al. postulated that the more similarity between a functional gene and its pseudogene, the higher the chance of pseudogene participating in homologous recombination causing gene conversion [15]. They provide an analogy between mutations in CYP21 gene, causing adrenal hyperplasia, to mutation in PKD1 gene. Both CYP21 gene and PKD1 share >95% of their sequence identity with their respective psuedogenes.
Distinguishing symptoms of PKD1 L845S mutation
All pathogenic mutations of ADPKD, whether frame-shift or missense or nonsense, have the same major symptom of renal cyst formation and associated kidney malfunction [2,13]. There are various other symptoms associated with the disease such as liver cysts, heart valve abnormalities (mitral valve regurgitation, mitral valve prolapse), hypertension, subarachnoidal aneurysm (SAH) or brain hemorrhage [2].
There are really no clear phenotypic characteristics that differentiate between all the known PKD mutations. However, there is a statistical data available on some nuances of phenotypes based on the study of a small population with closely related families. For example, in a study of PKD in Finnish population 11064 insC frame-shift mutation in PKD1 showed no mitral valve regurgitation or SAH, whereas L845S mutation had both these symptoms in 6/16 and 3/16 individuals [2]. The same study also collected data on age of the individual and onset of the disease, but found no correlation between the two. Also, there was no correlation found between missense, frame-shift or nonsense mutations and severity or age of disease onset. The only pattern that was reaffirmed was that mutations near the 5’ end of mRNA (nucleotides <7812) had early onset of ESRF (53 years vs. 56 years). ADPKD1 L845S mutation is on the extracellular part of PKD1 and has the diagnosis age of 37 ± 5 years and age onset of ESRF as 56 ± 3 years [2].
Transcriptional regulation
Transcription factors: Beta-catenin is a cell membrane protein that links actin cytoskeleton elements to cadherin family of trans-membrane cell to cell adhesion receptors. T-cell factors (TCF) are produced by T-cells and play a role in immune response. In the 3.3 kb promoter region near the 5’ end on PKD1, four TCF binding elements TBE (1-4) were identified [7]. The research was done on binding of beta-cateninn and TCF factors on TBE1. It was found that PKD1 is up-regulated and down-regulated by beta-cateninn and TCF factors respectively [7].
p53 is well known as tumor suppressor protein and 50% of cancer cases show a mutation of this gene [16]. It was found that in p53 deficient mice showed increased expression of PKD1 as compared to the wild type mice. Also, in human cells with DNA damage, which causes increased expression of p53, PKD1 expression was reduced. A model is proposed in which p53 represses the PKD1 promoter via recruitment of a histone deacetylase (HDAC) repressor complex [16]. HDAC represses transcription by preventing uncoiling of DNA around the nucleosomes. Hence, p53 is found to down-regulate PKD1 expression.
Retinoic acid (Vitamin A) derivatives, such as all-trans retinoic acid (AT-RA) and 9-cis retinoic acid (9C-RA) have been indirectly found to up regulate the PKD1 expression [17]. Vitamin A has a very important role in growth and differentiation of cells during embryonic development and growth of epithelial cells in adults. Implication of retinoic acid in controlling transcription of PKD1 is in line with trans-membrane location of PKD1 in epithelial cells. Retinoic acid derivatives do not directly bind to promoter regions of PKD1, but positively regulate it through general purpose transcription factor SP1 (Specificity Protein 1) [17].
Apart from SP1, there other common transcription factors that control PKD1 and PKD2 gene transcription such as E2F, EGRF, Ets, MZF1, Sp1, and ZBP-89 [18].
Alternative splicing
Alternatively splicing products of PKD1 mRNA to yield a functional protein is only documented in mouse brain but not kidney. This reported alternative splicing is between exons 12 and 13 of PKD1 gene [19]. Alternative splicing in human PKD1 was predicted from different cDNAs obtained for from nonunique mRNAs [3]. But, their source was not confirmed, perhaps due to pseudogenes that also get transcribed.
Post-translational modifications
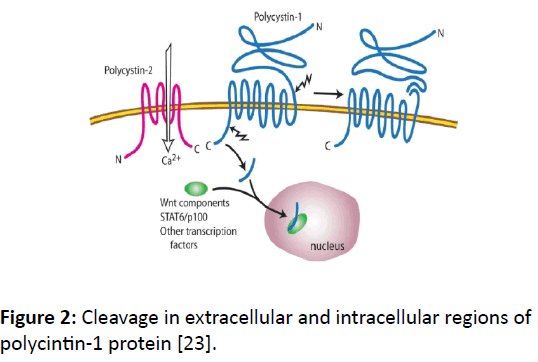
Cleavage: PKD1 is 450-kD protein and has two kinds of cleavage; one on the extracellular N-terminal GPS site and the other towards the cytoplasmic C-terminal tail (CTT) side (Figure 2). The N-terminal cleavage takes place at the GPS site, which separates the extracellular domains from the remaining domains in trans-membrane and intracellular regions. This GPS site autoproteolytic cleavage regulates kidney tubule development [20].
Figure 2: Cleavage in extracellular and intracellular regions of polycintin-1 protein [23].
There are two documented CTT cleavages triggered by low level of fluids in mouse kidney; one large ~35-kD [21] and the other 15-kD more proximal to C-terminal [22]. At least one of these N-terminal cleavages happens with association of PKD2. The cleaved portions of PKD2 move to the nucleus and bind with other transcription factors, such as STAT6 and p100 [22]. It is believed there are two forms of functional PKD1 protein, one fully intact and other cleaved.
Localization of PKD1: The tuberin protein, which is product of tuberous sclerosis 2 (TSC2) gene, is involved in the proper localization of PKD1 protein within the cell membrane [11]. PKD1 gene is only 63 base pairs downstream of TSC2 gene [24]. It was observed in infants, when heterozygous mutation were present in both TSC2 and PKD1 genes, ADPKD developed rapidly and with more severity as comptableared to the case when only PKD1 gene was mutated. In experiments with mice with defective TSC2, it was confirmed that PKD1 was found confined to golgi bodies, as opposed to the cell membrane in TSC2 normal mice. Golgi bodies are the sites of protein sorting. Administration of exogenous tuberin positioned the PKD1 protein to their correct location within the cell membrane [24]. This research confirmed that tuberin protein is involved in proper localization of PKD1 protein.
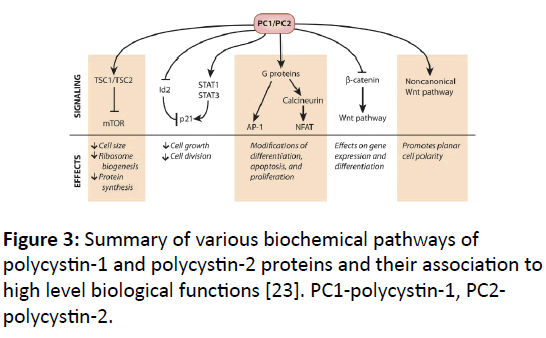
PKD1 function: A summary of PKD1 protein functions is presented in (Figure 3) and details are presented below.
Figure 3: Summary of various biochemical pathways of polycystin-1 and polycystin-2 proteins and their association to high level biological functions [23]. PC1-polycystin-1, PC2- polycystin-2.
Growth regulation: PKD1 and PKD2 exert negative control on excessive cell growth in kidney. This is evident from the uncontrolled development of cysts in defective PKD1 or PKD2 proteins. Both these proteins, directly or indirectly regulate the activity of other transcription factors or molecular signals that cause enhanced cell growth. PKD1 is an inhibitor of mTOR (mammalian target for rapamycin) pathway [25]. mTOR causes increased cell growth and it is inhibited by a complex of TSC1 and TSC2 (tuberous sclerosis 1 and 2). mTOR is activated my mTOR kinase, which requires another protein called Rheb to be in its GTP bound state. The complex of TSC1 and TSC2, which is stabilized by PKD1, acts as a GTPase (hydrolyzes GTP) of Rheb that down regulates mTOR kinase.
Cell cycle is regulated by cyclin dependent kinases (Cdks), which is inhibited by p21. PKD1 and PKD2 work together to elevate the levels of p21. PKD1 can increase the levels of p21 by binding to Janus Kinase (JAK) and members of signal transducers and activators of transcription (STAT) pathway [26]. PKD1 activates STAT1 and STAT3 and increases levels of p21.
G-Protein activation: C-terminal tail (CTT) end of PKD1 activates G-protein, which has 3 domains (heterotrimeric). Activated G-Protein alpha subunits interact with c-Jun Terminal kinase (JNK) and transcription factor AP-1 to positively regulate their activity [27]. AP-1 is associated with apoptosis, cell differentiation and multiplication through many different signaling pathways.
PKD1 also activates nuclear factor of activated T-cells (NFAT) [28]. NFAT regulates genes involved in apoptosis, growth, differentiation and cell adaption. Artificially controlled expression of PKD1 causes NFAT accumulation in nucleus, and this effect is enhanced by co-expressing Gaq, a known PKD1- binding G protein subunit [28]. NFAT is also connected to calcium signaling and PKD2 localization. NFAT is activated by calcineurin, which is activated by increased Ca2+ levels in the cytosol. Calcineurin dephosphorylates NFAT which leads to its nuclear accumulation. Phosphorylation of NFAT by Glycogen synthase kinase 3-beta (GSK 3-beta) causes NFAT to move back into the cytoplasm. Calcineurin is up regulated by G-protein activated through PKD1 [28].
Wnt signaling: Wnt pathway consists of a set of proteins involved in growth, development and planar cell polarity. PKD1 negatively regulates Wnt signaling through beta-catenin dependent (canonical) and beta-catenin independent (noncanonical) pathways [29]. In the canonical pathway Wnt cell surface receptor accepts beta-catenin and facilitates its movement to the nucleus and activates T cell factor (TCF) transcriptional activity. TCF is a general purpose transcription factor. Cleaved CTT end of PKD1 binds to beta-catenin, moving with it to the nucleus and negatively regulating TCF dependent transcription.
Wnt non-canonical signaling is related to cell polarity by making sure the cells near the kidney tubule divide parallel to the axis of the tube [23]. PKD1 is believed to regulate this noncanonical Wnt signaling. In ADPKD, the cells are seen to differentiate at an angle rather than parallel to the tubule axis, which leads to uneven rather than tube like growth in kidney tubules.
Regulation
Actin cytoskeleton: PKD1 protein exists as a complex with other proteins in the cell membranes of different cell types. It exists with integrins to form cell-matrix (focal) adhesion complexes or with E-cadherin-beta-catenin complex to form cell to cell adhesion complexes [13]. Both these types of complexes associate with actin cytoskeleton through the cell membrane. This association has lead researchers to study the role of actin cytoskeleton and PKD1 protein complexes. They found on using Cytochalasin D., which depolymerizes actin microfilament, both focal adhesion and cell to cell adhesion complexes with PKD1 were completely disrupted [11]. This research finding has demonstrated the control of PKD1 containing complexes by actin cytoskeleton.
Phosphorylation: Phosphorylation of polycystin-1 protein in Human Fetal Collecting Tubule (HFCT) cells was demonstrated by precipitating PKD1 protein out using its antibody and then doing a Western blot analysis with antiphosphotyrosine (APHT) antibody. The APHT antibody is used to extract phosphorylated tyrosines so their amount can be measured. The phosphorylation of tyrosine is widely known in various signal transduction and cell to cell communication pathways. In cells with ADPKD reduced levels of tyrosine phosphorylation was found, which implicates tyrosine phosphorylation for polycystin-1 regulation [11].
Further research with protein kinase a (PKA) and c-src tyrosine kinase have found some specific sites of tyrosine, serine/threonine phosphorylation near the C-terminal of PKD1 protein [30]. PKA, also known as cyclic AMP (cAMP) dependent kinase, activates serine/threonine sites and c-src activates tyrosine sites through phosphorylation. These findings affirm the roles of phosphorylation on polycystin-1 protein control.
Calcium: Polycystin-1 contains extracellular C-lectin and REJ domain related to Ca2+ ion related binding or influx. More convincing evidence of role of calcium is found in HFCT cells in regulation of polycystin-1 protein complexes found in cell-matrix (focal adhesion) and cell-to-cell junctions [11]. It is found that increased calcium concentrations lead to increase in association of polycystin-1-E-cadherin-beta catenin complex (cell to cell adhesion) and decreased association of focal adhesion kinase (FAK)-polycystin-1 (cell-matrix) associations [31]. The cell-matrix junctions are predominant during early development and migration of cells in ureteric bud and later in adult cell-to-cell junctions are predominant in fully differentiated in renal collecting tubules [11].
miRNA: MicroRNAs (miRNAs) are short, about 22 nucleotides long, non-coding RNAs. The important regulatory roles played by miRNAs have only been highlighted in recent years that include gene regulation, cell proliferation, differentiation and apoptosis. The first evidence of miRNAs in ADPKD was provided by Lee et al. by analyzing epithelial cells of bile ducts (cholangiocytes) from rats with polycystic kidneys [32]. A miRNA, named miR-15a, was identified whose expression in cholangiocytes cell line lines (CCL) of polycystic rats was found to be 37 times lower than normal control rats and also significantly lower in liver cells of human ADPKD or ARPKD patients [32]. Reduction in miR-15a levels causes up-regulation of a cell cycle regulator protein Cdc25A. Laboratory controlled up regulation of mi-R15a in rat PKD-CCL caused decreased Cdc25A levels and decreased cell proliferation, while down regulation of mi-R15a in normal rat increase cell proliferation and cyst growth. As regards the direct role of miRNA in controlling gene expression of PKD1 gene, several sites have been found in the 3’ UTR region of both PKD1 and PKD2 that are believed to be the target of these miRNAs: miR-200b/c, miR-429, miR17-5p, miR-20 etc. [4].
Disease characteristics
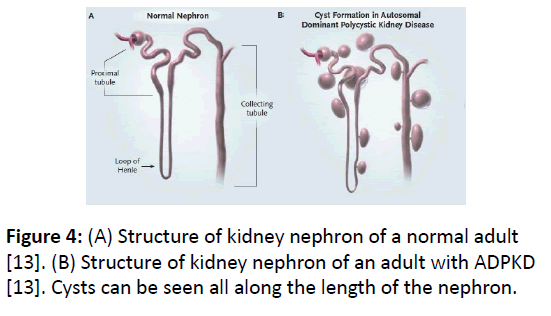
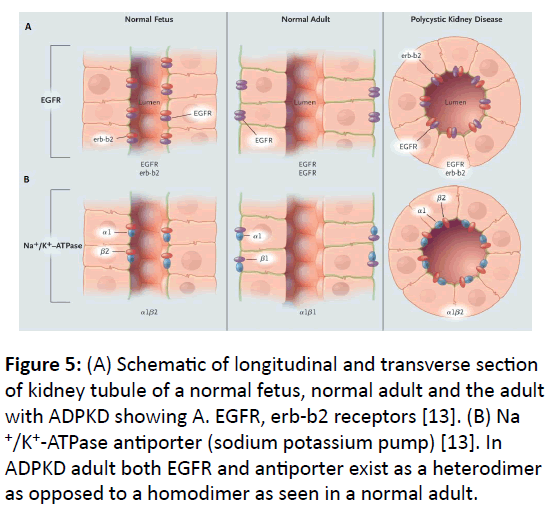
Cyst development: Normal development of kidney tubule is programmed by precise control between cell proliferation and cell death (apoptosis). In ADPKD1, this control is disturbed in favor of persistent apoptosis due to defective protein product of mutated PKD1 gene, which causes development of cysts all along the length of nephron (Figure 4). In normal individuals, the proliferation of epithelial cells of kidney tubules ceases after birth but in ADPKD, epithelial cells develop into cysts throughout the adult life. Over-expression of epidermal growth factor (EGFR) in ADPKD, which is present in the epithelial cells of kidney tubule epithelia, is implicated in the development of cysts [13]. EGFR exists as a complex of dimers of two same units in adults (homodimer), but as heterodimer with its variant erbb2 in normal fetus or polycystic kidney as shown in (Figure 5A). Also, the location of EGFRs and antiporters in ADPKD adult is different than the normal adult.
Figure 5: (A) Schematic of longitudinal and transverse section of kidney tubule of a normal fetus, normal adult and the adult with ADPKD showing A. EGFR, erb-b2 receptors [13]. (B) Na +/K+-ATPase antiporter (sodium potassium pump) [13]. In ADPKD adult both EGFR and antiporter exist as a heterodimer as opposed to a homodimer as seen in a normal adult.
Secretion: In normal kidney, proper filtration balance of Na+, K+ and Cl- is achieved by the sodium-potassium pump (Na+/K+/ ATPase antiporter) and Na+, K+, 2Cl- symporter [5]. In the ADPKD adult the location of these pumps is different than that of a normal adult as shown in (Figure 5B), in which the antiporter is located in the lumen of the tubule rather than towards the basolateral region. Also, the antiporter in ADPKD consists of a heterodimer of alpha-1 and beta-2 subunits as opposed to a homodimer of alpha-1 and beta-1 subunits [13] (Figure 5B). These defects of antiporters and symporters cause kidney malfunction in disturbing the proper balance of Na+, K+ and Clfiltration. The cysts in ADPKD secrete increased amount Cl- ions in the cyst lumen, stimulated by cAMP, involving transmembrane protein known as cystic fibrosis trans-membrane regulator (CFTR) located in cyst epithelial cells [33].
Diagnosis
In individuals with a family history of ADPKD, a combination of age based renal imaging technologies, such as CTscan, MRI or ultrasonography, can be employed to determine the presence or status of progression of the disease [4]. In individuals with no family history genetic testing can be useful in prognosis and diagnosis of the disease. Two methods employed for genetic testing are: DNA linkage analysis and gene based mutation screening.
Linkage analysis is performed using some specific microsatellite markers (e.g., short tandem repeats) near the vicinity of PKD1 or PKD2 genes and is suitable only in cases where many family members are either unwilling or not available to participate in the study. After construction of linkage map for the two ADPKD genes in a homologous chromosome pair, a haplotype analysis can be performed to predict the disease status of other individuals of the family [4]. This method is used very rarely, but can be used in prenatal testing, when the mutation type in two ADPKD genes is unknown.
Gene based mutation screen can be performed either DNA or mRNA. A full gene analysis strategy and mRNA based protein truncation test (PTT) was proposed by Rossetti et al [34]. They suggested an approach by using long range polymerase chain reaction (LR-PCR) to amplify the duplicated sequences in PKD1 domains from exon 1 to exon 33. The approach Rossetti et al. suggested overcame the previously difficulties in getting copies of duplicated region of PKD1 gene, due to interference of several non-coding psuedogenes that have similar nucleotide sequences as the real coding PKD1 gene. The efficacy of full gene analysis based genetic testing of ADPKD was studied by Garcia et al. [35]. They were able to detect a broad range of mutations using the full gene sequence analysis; missense mutations, in frame insertion/deletion and non-customary splicing mutations.
Treatment
There is no single treatment to replace the malfunctioning polycystin proteins, but there are strategies to mitigate the two key disturbances which are conspicuous in the slow progression of the disease; proliferation of epithelial cells leading to cyst expansion, abnormal secretions that release rather than absorb fluids and electrolytes.
Patricia et al. propose the development of EGFR inhibitors to retard the progression of the cysts [13]. The above normal mTOR activity in cysts formation lead researchers to study treatment avenue of effect of down-regulating mTOR activity [26]. However, with mice experiments, there were several side effects related to limiting mTOR activity. For example, mTOR inhibitors restrict nitric oxide synthesis (a natural neurotransmitter), restrict survival of endothelial cells, increase the risk of capillary loss in kidney and increase hypertension [36].
Due to increased secretion of Cl- ions stimulated by cAMP in CTFR chloride channel in cysts, down regulation of cAMP and inhibition of CTFR chloride channels are considered as possible molecular targets [37,33]. The CFTR inhibitor compound CFTRinh172 showed dramatic reduction in cyst growth [37]. Antidiuretic hormone (ADH), also known as vasopressin, is a major up-regulator of cAMP production through V2 vasopressin receptors located in kidney collecting tubules. A drug named Tolvaptan, which inhibits the V2 receptors, is found to significantly reduce cyst growth in mice [38]. Tolvaptan was recently approved by the FDA as the first drug specifically designed to treat polycystic kidney disease [39]. Bardoxolone methyl is in clinical trials for patients with diabetes and chronic kidney disease from PKD [39,40]. The main mechanism of action of bardoxolone is the activation of Nrf2, a transcription factor involved in the up-regulation of many cytoprotective genes with consecutive anti-inflammatory effects, inhibition of nuclear factor κB (NF-κB), and decreased oxidative stress [40].
Discussion and Conclusion
There exist a large number of mutations in PKD1 gene that cause ADPKD1, especially in the duplicated domains of the gene. They can be frame-shift single insertion/deletion, missense or a large deletion. Due to the large size of the gene, many of these mutations are private and may not be widespread in other populations. There are also large number non-coding pseudogenes of PKD1, which may also attribute to mutation due to gene conversion events. The existence of pseudogenes also made PCR based analysis of the disease mutation difficult, because the primers of PCR would bind to psuedogenes that have homologous nucleotide sequences in comparison to the real coding PKD1 gene.
In ADPKD1, the location and sequence of causative genes have been identified, but the comprehensive functions of the protein product polycystin-1 have yet to be ascertained. This is again due to large size of the gene that leads to the complexity of polycystin-1. Polycystin-1 works in association with a related protein polycystin-2 and with other integral membrane proteins in multi-protein complexes. These multi-protein complexes are involved in various cell-to-cell communication and intracellular signal transduction pathways including many gene transcription regulating activities. Unless these functions, however complex and difficult to grasp, are properly understood, in both normal and diseased individuals, satisfactory treatment of the ADPKD1 disease will remain an elusive goal.
Future Directions
One of the intriguing aspects regarding the polycystin-1 protein is its function in association with primary (singular) cilia present in kidney tubule epithelial cells [3,4]. Even though there are some cases where the structure of these cilia was malformed in kidney tubules of polycystic mouse, not much is known about these cilia, let alone the association of polycystin-1 with them [41].
Although recent research have demonstrated role of miRNA in regulation of ADPKD genes, several questions remain unanswered regarding their putative significant role in embryonic kidney development or even in adult kidneys; whether they work in regulation at transcription or translation level, possible role of introns as coding sequences for premiRNAs and mutations in such introns [4]. For example, there is research on predicted role of an endogenous generic miRNA miR-200c on possible degradation of PKD1, but there is no in vivo demonstration that proves or disproves this role of miR-200c and also whether it acts in degrading the mRNA transcript of PKD1 or prevents its translation by attaching to mRNA 3’ UTR region [4]. Within intron 45 of PKD1 a miRNA gene is found. This miRNA name is miR-1225 and it belongs to the Mirtron family [40-44]. Mirtrons are introns that code for miRNA. Even though polycystic disease causing mutations have been reported in splicing of exon 45, not much is known about how miRNA miR-1225 works or how a mutation in this miRNA causes polycystic kidney disease.
The avenue of gene therapy for treatment of ADPKD to fix the defective PKD1 or PKD2 genes can be explored further. Some research in use of gene therapy have been shown to be of not much use after cysts have already developed, since a broad range of kidney tissues are impacted [10]. But, there is no gene therapy treatment research at a very early age, say infancy, after positive identification from genetic tests. Given ADPKD shows up in mid-life on average, early diagnosis and treatment using an emerging technology like gene therapy may have promising results.
References
- Somlo S (1998) Polycystic Kidney Disease Genes and Polycystins. ClinExpNephrol 2: 211-217.
- Peltola P, Lumiaho A, Miettinen R, Pihlajamaeki J, Sandford R, et al. (2005) Genetics and phenotypic characteristics of autosomal dominant polycystic kidney disease in Finns. J Mol Med 83: 638-646.
- Polycystic kidney disease: the complete structure of the PKD1 gene and its protein. The International Polycystic Kidney Disease Consortium. Cell 81: 289-298.
- Tan YC, Blumenfeld J, Rennert H (2011) Autosomal dominant polycystic kidney disease: Genetics, mutations and microRNAs. Biochimica et BiophysicaActa 1812: 1202-1212.
- Thomas RL, McConnell R, Whittacker J, Kirkpatrick P, Bradley J, et al. (1999) Identification of Mutations in the Repeated Part of the Autosomal Dominant Polycystic Kidney Disease Type 1 Gene (PKD1) by Long-Range PCR. Am J Hum Genet 65: 39-49.
- Chapin HC, Caplan MJ (2010) The cell biology of polycystic kidney disease. J Cell Biol 191: 701-710.
- Rodova M, Islam MR, Maser RL, Calvet JP (2002) The Polycystic Kidney Disease-1 Promoter Is a Target of the beta-Catenin/T-cell Factor Pathway. J BiolChem 277.
- Chauvet V, Qian F, Boute N, Cai Y, Phakdeekitacharoen B (2002) Expression of PKD1 and PKD2 transcripts and proteins in human embryo and during normal kidney development. Am J Pathol 160: 973-983.
- Bycroft M, Bateman A, Clarke J, Hamill SJ, Sandford R, et al. (1999) The structure of a PKD domain from polycystin-1: implications for polycystic kidney disease. EMBO J 18: 297-305.
- Kleymenova E, Ibraghimov-Beskrovnaya O, Kugoh H, Everitt J, Xu H, et al. (2001) Tuberin-dependent membrane localization of polycystin-1: a functional link between polycystic kidney disease and the TSC2 tumor suppressor gene. Mol Cell 7: 823-832.
- The PKD Mutation Database Version 2.1. Autosomal Dominant Polycystic Kidney Disease: Mutation Database (2011). Polycystin-1 precursor – Homo Sapiens. UniProtKB.
- Polycystic kidney disease 1 (Autosomal Dominant). Genetics Home Ref.
- Wilson PD (2004) Polycystic Kidney Disease. N Engl J Med 350: 151-164.
- Wilson PD (2001) Polycystin: New Aspects of Structure, Function, and Regulation. J Am SocNephrol 12: 834-845.
- Moy GW, Mendoza LM, Schulz JR, Swanson WJ, Glabe CG, et al. (1996) The sea urchin sperm receptor for egg jelly is modular protein with extensive homology to the human polycystic kidney disease protein PKD1. J Cell Biol 133, 809-817.
- Watnick TJ, Gandolph MA, Weber H, Neumann HPH, Germino GG (1998) Gene conversion is a likely cause of mutation in PKD1. Hum Mol Genet 7: 1239-1243.
- Bodegom DV, Saifudeen Z, Dipp S, Puri S, Magenheimer BS, et al. (2006) The Polycystic Kidney Disease-1 Gene Is a Target for p53-mediated Transcriptional Repression. J BiolChem 281: 31234-31244.
- Islam MR, Puri S, Rodova M, Magenheimer BS, Maser RL, et al. (2008) Retinoic acid-dependent activation of the polycystic kidney disease-1 (PKD1) promoter. Am J Physiol Renal Physiol 295: F1845-F1854.
- Lantinga-van Leeuwen IS, Leonhard WN, Dauwerse H, Baelde HJ, van Oost BA, et al. (2005) Common regulatory elements in the polycystic kidney disease 1 and 2 promoter regions. Eur J Hum Genet 13: 649-659.
- Lohning C, Nowicka U, Frischauf AM (1996) The mouse homolog of PKDI: sequence analysis and alternative splicing. Mamm Genome 8: 307-311.
- Yu S, Hackmann K, Gao J, He X, Piontek K, et al. (2007) Essential role of cleavage of Polycystin-1 at G protein-coupled receptor proteolytic site for kidney tubular structure. ProcNatlAcadSci USA. 104: 18688-18693.
- Chauvet V, Tian X, Husson H, Grimm DH, Wang T, et al. (2004) Mechanical stimuli induce cleavage and nuclear translocation of the polycystin-1 C terminus. J Clin Invest 114: 1433-1443.
- Low SH, Vasanth S, Larson CH, Mukherjee S, Sharma N, et al. (2006) Polycystin-1, STAT6, and P100 Function in a Pathway that Transduces CiliaryMechanosensation and Is Activated in Polycystic Kidney Disease. Develop Cell 10: 57-69.
- Chapin HC, Caplan MJ (2010) The cell biology of polycystic kidney disease. J Cell Biol 191: 701-710.
- Hughes J, Ward CJ, Peral B, Aspinwall R, Clark K, et al. (1995) The polycystic kidney disease 1 (PKD1) gene encodes a novel protein with multiple cell recognition domains. Nat Genet 10: 151-160.
- Shillingford JM, Murcia NS, Larson CH, Low SH, Hedgepeth R, et al. (2006) The mTOR pathway is regulated by polycystin-1, and its inhibition reverses renal cystogenesis in polycystic kidney disease. ProcNatlAcadSci USA 103: 5466-5471.
- Bhunia AK, Piontek K, Boletta A, Liu L, Qian F, et al. (2002) PKD1 induces p21(waf1) and regulation of the cell cycle via direct activation of the JAK-STAT signaling pathway in a process requiring PKD2. Cell 109: 157-168.
- Arnould T, Kim E, Tsiokas L, Jochimsen F, Grüning W, et al. (1998) The polycystic kidney disease 1 gene product mediates protein kinase C alpha-dependent and c-Jun N-terminal kinase-dependent activation of the transcription factor AP-1. J BiolChem 273: 6013-6018.
- Puri S, Magenheimer BS, Maser RL, Ryan EM, Zien CA, et al. (2004) Polycystin-1 activates the calcineurin/NFAT (nuclear factor of activated T-cells) signaling pathway. J BiolChem 279: 55455-55464.
- Lal M, Song X, Pluznick JL, Di GV, Merrick DM, et al. (2008) Polycystin-1 C-terminal tail associates with beta-catenin and inhibits canonical Wnt signaling. Hum Mol Genet 17: 3105-3117.
- Li HP, Geng L, Burrow CR, Wilson PD (1999) Identification of phosphorylation sites in the PKD1-encoded protein C-terminal domain. BiochemBiophys Res Commun 259: 356-363.
- Geng L, Burrow CR, Li HP, Wilson PD (2000) Modification of the composition of polycystin-1 multiprotein complexes by calcium and tyrosine phosphorylation. BiochimBiophysActa 1535: 21-35.
- Lee SO, Masyuk T, Splinter P, Banales JM, Masyuk A, et al. (2008) MicroRNA15a modulates expression of the cell-cycle regulator Cdc25A and affects hepatic cystogenesis in a rat model of polycystic kidney disease. J Clin Invest 118: 3714-3724.
- Davidow CJ, Maser RL, Rome LA, Calvet JP, Grantham JJ (1996) The cystic fibrosis trans-membrane conductance regulator mediates transepithelial fluid secretion by human autosomal dominant polycystic kidney disease epithelium in vitro. Kidney Int 50: 208-218.
- Rossetti S, Strmecki L, Gamble V, Burton S, Sneddon V, Peral B, et al. (2001) Mutation analysis of the entire PKD1 gene: genetic and diagnostic implications. Am J Hum Genet 68: 46-63.
- Garcia-Gonzalez MA, Jones JG, Allen SK, Palatucci CM, Batish SD, et al. (2007) Evaluating the clinical utility of a molecular genetic test for polycystic kidney disease. Mol Genet Metab 92: 160-167.
- Torres VE, Boletta A, Chapman A, Gattone V, Pei Y, et al. (2010) Prospects for mTOR inhibitor use in patients with polycystic kidney disease and hamartomatous diseases. Clin J Am SocNephrol 5: 1312-1329.
- Yang B, Sonawane ND, Zhao D, Somlo S, Verkman AS (2008) Small molecule CFTR inhibitors slow cyst growth in polycystic kidney disease. J Am SocNephrol 19: 1300-1310.
- Gattone VH, Wang X, Harris PC, Torres VE (2003) Inhibition of renal cystic disease development and progression by a vasopressin V2 receptor antagonist. Nat Med 9: 1323-1326.
- Steinman T (2018) Beyond tolvaptan, what lies ahead: New therapies in the treatment of polycystic kidney disease. Nephrol News Issues.
- Dick Z, Akizawa T, Audhya P, Bakris GL, Chin M, et al. (2013) Bardoxolone Methyl in Type 2 Diabetes and Stage 4 Chronic Kidney Disease. N Engl J Med 369: 2492-2503.
- Yoder BK, Hou X, Guay-Woodford LM (2002) The polycystic kidney disease proteins, polycystin-1, polycystin-2, polaris and cystin, are co-localized in renal cilia. J Am SocNephrol 13: 2508-2516.
- The PKD Mutation Database Version 2.1. Autosomal Dominant Polycystic Kidney Disease: Mutation Database (2011). Polycystin-1 precursor–Homo Sapiens. UniProtKB.
- Imai E, Isaka Y (2004) Perspectives for gene therapy in renal diseases. Intern Med 43: 85-96.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences